The CTD² Network and Cancer Systems Biology Consortium organized a virtual symposium series titled “Multidisciplinary Approaches to Understand Cancer Treatment Resistance”. Please join us on 11/16, 11/17, 12/2, 12/16, and 12/17. Click here to view the registration website.
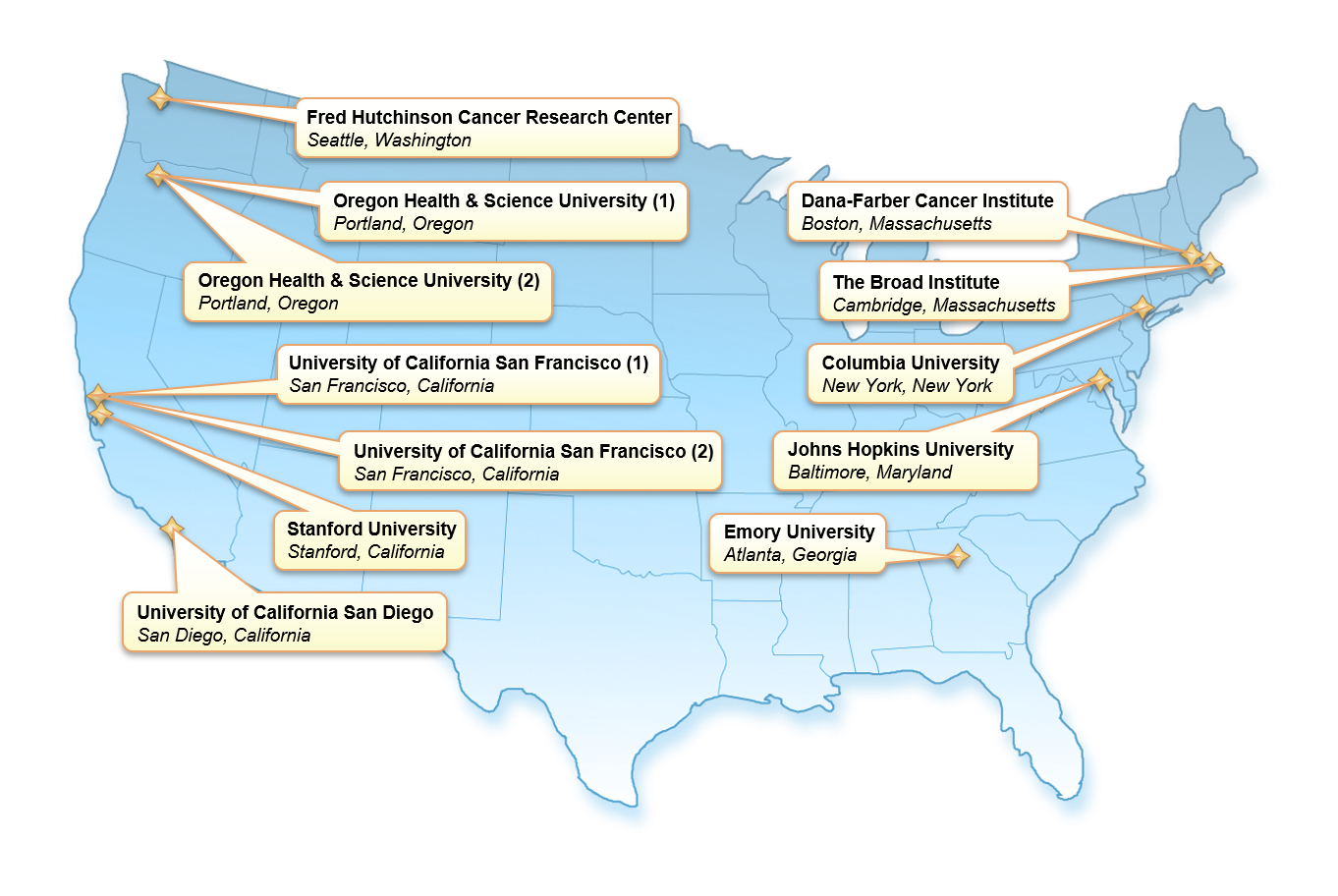
CTD² Centers
The CTD2 Network focuses on identifying and understanding (1) pathways that influence cancer phenotypes (including understanding the function of the genes/target that are essential during cancer initiation, progression and maintenance); (2) perturbagens, singly or in combination, that can modulate such pathways; and (3) biomarkers which predict responses to treatments, help determine prognosis, and/or contribute to understanding of other aspects of cancer etiology to enable the development of effective therapeutic modalities in the future. The Network also addresses whether these findings are dependent on the defined genetic background (either germline or somatic) of the patient.
CTD2 Network is comprised of 12 OCG-supported research teams, called Centers, led by a principal investigator. The Centers work independently as well as collaboratively as a “Network”. Network Centers utilize a combination of state-of-the-art high throughput informatic and experimental approaches to functionally validate discoveries from genomic studies and advance them toward precision oncology. They share data and resources, such as analytical tools and reagents, across the Network and with the research community.
Each Center contributes unique expertise in one or more of the following areas:
- In-depth mining of genomic data
- Systems biology analyses
- High-throughput screening of small molecules or genetic perturbagens
- In vitro and in vivo biological characterization approaches
- Cancer models (cell lines, organoids, PDXs)
- Gene-gene dependencies (e.g., synthetic lethality)
- Identification of interacting protein subunits in biochemical processes
- Complex analytical approaches to target identification
- Technology development and improvement
The following descriptions inform what each Center is contributing to the Network and the greater research community.
Current Phase
Targeting Vulnerabilities of Therapy-resistant Cancer Cell States with Small Molecules
Principal Investigator: Stuart L. Schreiber, Ph.D.
Targeted therapies and immunotherapies (immune checkpoint inhibitors) have been the most transformative advances in cancer treatment in the last several years. However, these therapies, like traditional chemotherapies, eventually result in tumor resistance to therapeutic attacks on their vulnerabilities. Hence, understanding how to avoid or overcome resistance to therapies is essential when developing treatments that are effective over the long term. The CTD2 Center at the Broad Institute aims to discover and understand the bases of therapy resistant-state vulnerabilities and exploit these susceptibilities to develop effective combination therapies.
In the preceding phase, the Broad Institute developed powerful new tools and capabilities that enable the community to identify novel cancer vulnerabilities. These capabilities are shared using the Cancer Therapeutics Response Portal (CTRP), which houses a large dataset of compound sensitivity data (concentration-response curves) and has been made available without restriction. Using the CTRP, researchers at this Center found evidence for the existence of at least one common therapy-resistant state associated with mesenchymal characteristics of cancer cells. This state emerges by non-mutational mechanisms after treatment with either chemotherapy or targeted therapeutics across several cancer types. In the current phase, the Center proposes to i. discover common drug-resistant cancer cell states, ii. define vulnerabilities of these cancer cell states, and iii. identify small molecules that can attack these susceptibilities. These studies will be explored in the context of patient-derived cancer models, which will provide a path forward for combination therapies to overcome treatment-resistance.
Systematic Identification and Pharmacological Targeting in Tumor Dependencies for Precision Cancer Medicine
Principal Investigator: Andrea Califano, Ph.D.
Cancer is a complex and highly heterogenous disease, hence an individual gene/protein may not represent an effective target for all the sub-clone-specific mutations in a tumor. Indeed, while genetic-based targeted therapy and immunoncology hold great promise, most patients still do not respond, or will eventually relapse with drug resistant tumors, suggesting that the concept of therapeutic targets as single proteins may need to be revisited. Earlier studies revealed that tumor cell state depends on the coordinated activity of a handful of aberrant master regulator (MR) proteins. These proteins form small hyperconnected components termed tumor checkpoint modules (TCM), which are tightly regulated. The aberrant activity of a TCM is induced by genetic or epigenetic (genomic) alterations in upstream pathways. The CTD2 Center at Columbia aims to use this TCM/MR-based conceptual framework to elucidate new druggable targets for single agent and combination therapy.
In the preceding phase, Columbia University has shown that comprehensive dissection of tumor-specific gene regulatory layers can help elucidate novel tumor dependency mechanisms. In the current phase, the Center proposes to i. perform network-based analyses of tumor samples (atypical/anaplastic meningioma, SDHBDel gastrointestinal stromal tumors, and metastatic bladder, lung, and colon adenocarcinoma), ii. prioritize and characterize FDA-approved and late-stage investigational drugs by assessing differential tumor checkpoint activity, and iii. evaluate these single and combination targets in patient-derived xenograft and organoid models. These studies provide a mechanistic approach for precision oncology, where therapeutic targets, associated small molecule inhibitors, and population stratification biomarkers are derived from rigorous and detailed understanding of tumor state regulation and drug-induced modulation.
The Dana-Farber Cancer Institute Cancer Target Discovery and Development Center
Principal Investigator: William C. Hahn, M.D., Ph.D.
Efforts to characterize cancer genomes provide a view of the mutations and copy number alterations that occur in human cancers. However, it remains unclear which of these alterations are critical for tumor maintenance, heterogeneity, and the ability to evade the immune system. Identifying genes that are essential for tumor survival and immune evasion will accelerate the development of new molecularly targeted therapeutics. The emerging clinical success of checkpoint blockade is tempered by the reality that most patients do not respond to immunotherapy. New targets are needed to improve tumor responses and guide rational combination immunotherapy to overcome resistance. The CTD2 Center at the Dana-Farber Cancer Institute (DFCI) aims to define a comprehensive classification of targets, develop means to rationally identify combination therapies, and discover genes that modulate the response to immunotherapeutics.
In the preceding phase, DFCI developed the bioinformatic tools, methods, and infrastructure to perform genome-scale loss-of-function and gain-of-function screens in human cancer cell lines and patient-derived models to identify and validate cancer targets. In the current phase, the Center proposes to i. systematically investigate genes essential for cancer cell proliferation and survival in colon and pancreatic cancers, ii. interrogate the effects of manipulating combinations of genes and targets to understand the pathways that program malignant transformation, and iii. identify cancer driver genes that enable tumor cells to evade the immune system to uncover new immunotherapy targets and develop rational combinations. These studies will provide functional insights that complement genomic data to identify potential targets and facilitate the translation of this information into the development of therapeutics and diagnostics. The data and methodological approaches used by this Center will be readily available to the CTD2 Network and scientific community.
Systematic Discovery of Neomorph Protein-Protein Interactions in Cancer for Oncogenic Pathway Perturbation
Principal Investigator: Haian Fu, Ph.D.
Genomic mutational landscape offers a unique opportunity to distinguish cancer cells from normal cells, with re-wired oncogenic pathways and networks for therapeutic interrogation. Current cancer genomics efforts are largely centered on identifying cancer-causing genes as therapeutic targets and biomarkers. However, a large number of gene mutations give proteins new capabilities to bind cellular proteins and create new signaling pathways that drive tumor growth. Hence, understanding how to leverage these genomic changes at the mutated amino acid resolution for cancer-specific target discovery, and how to rapidly translate this knowledge into genotype-directed cancer therapies for precision oncology, remains a daunting and urgent challenge. The CTD2 Center at Emory University aims to understand the functions of genomic mutations in cancer etiology through systematic interrogation of protein-protein interactions (PPIs) for target identification, validation, and therapeutic discovery across solid and hematological cancer types.
In the preceding phase, the Center developed a sensitive live cell-based biosensor platform coupled with bioinformatics pipeline for exploring PPIs in a high throughput format. This quantitative High Throughput differential Screening platform, termed qHT-dS, enables comparative screening of wild type and mutant driver genes. This platform can be used to discover mutant allele-specific gain-of-function (neomorph) oncogenic PPIs (neo-PPIs). In the current phase, Emory University’s CTD2 Center aims to leverage rich genomic mutation data to discover and validate cancer-specific PPIs as targets and pathway perturbagens to accelerate patient-centered therapies. To accomplish this, the Center proposes to i. identify and validate cancer mutation-created neo-PPIs through differential screening with the qHT-dS technology platform, ii. identify small-molecule perturbagens of selected neo-PPIs and understand the drug resistance mechanism, and iii. improve and implement informatics pipeline for streamlined data analysis and integration of genomics information with neoPPI-mediated oncogenic networks and therapeutic responses. These studies will provide potential molecular mechanisms and reveal promising cancer-specific targets for genotype-directed therapeutic discovery.
Personalized Cancer Models to Discover and Develop New Therapeutic Targets
Principal Investigator: Christopher Kemp, Ph.D.
A major goal of personalized oncology is to use a tumor’s DNA sequence, gene expression, or other molecular data to inform patient care. Large-scale molecular characterization studies now provide a comprehensive view of the genomic landscape of most human cancers. However, many commonly mutated cancer genes are difficult to target with drugs, and even for genes that might be targetable, it is not clear which one should be prioritized or which drug would be effective for any given patient. Identifying actionable gene targets that are clinically effective in the context of genotypic and phenotypic heterogeneity of tumors remains a major challenge. Moreover, even in cases where targeted therapy works, development of resistance is common, which further highlights the need for additional targeted agents and effective drug combinations. To address these challenges, the CTD2 Center based at the Fred Hutchinson Cancer Research Center (FHCRC) developed an approach whose main innovation is functional genomic and drug profiling of patient-derived tumor cells to identify gene targets and candidate drugs with associated biomarkers for preclinical development.
In the preceding phase, the FHCRC Center developed and optimized a pharmaceutical industry grade, array-based high-throughput siRNA and drug screening platform which accurately and efficiently identifies tumor- and genotype-specific vulnerabilities. In the current phase, the Center proposes to i. identify cancer- and genotype-specific therapeutic targets (e.g. synthetic lethal genes) and effective drug combinations to overcome drug resistance using both isogenic and patient-derived tumor cell cultures, ii. develop novel computational approaches and informatics tools to prioritize gene targets with associated biomarkers using large scale data sets, and iii. validate prioritized targets with orthogonal assays in genomically characterized patient-relevant tumor models of increasing complexity and heterogeneity. The Center will initially focus on pancreatic and ovarian cancer with capabilities to extend to additional cancers. These studies will nominate novel targets suitable for drug development and propose more effective therapeutic strategies for several cancer types, particularly for advanced and chemotherapy-resistant tumors.
Pathway Discovery and Target Validation for Outgrowth of Breast Cancer Metastasis
Principal Investigators: Joel S. Bader, Ph.D. and Andrew Ewald, Ph.D.
Majority of cancer deaths are attributable to metastasis, rather than growth of the primary tumor. Tumors are heterogeneous at the genetic, epigenetic, and phenotypic levels; thereby effective cancer treatments are limited. Current diagnostic approaches do not allow individualized assessment of metastatic recurrence risk nor do they allow effective therapies for metastatic cancers. Breast cancers have few common mutations, and each tumor seems a unique mixture of different low frequency mutations. In breast cancer, metastatic recurrence may occur years to decades after treatment; hence, breast cancer presents a unique research opportunity to improve patient outcomes if effective anti-metastatic therapies could be developed. The CTD2 Center at Johns Hopkins University (JHU) aims to combine advances in experimental (Ewald) and computational (Bader) methods to interrogate the metastatic process and systematically dissect the genetic basis of breast cancer.
This Center has developed and applied computational methods to connect quantitative features to their genetic basis across multiple complex human diseases. The Center aims to use a pipeline that relies on organoids from primary human breast cancer tissue to model several distinct steps of metastasis. The Center plans to combine organoids developed from primary human breast tumors with real-time imaging to convert the process of cancer progression into quantitative phenotypes that can be dissected systematically by genomics technologies. To accomplish this goal, the Center proposes to i. identify complex features (or molecular correlates) in primary human breast tumor organoids required for the initial metastatic process, ii. utilize network analysis techniques to prioritize and validate if these candidate targets are required for metastatic growth in the organoid system and patient-derived xenograft models, and iii. modulate candidate targets with chemical and genetic perturbagens from the CTD2 Network and other drug discovery efforts. These studies will identify actionable targets for preventing metastatic recurrence or treating patients with established breast cancer metastases.
Functional Genomic Discovery of Pathway Targeted and Immune Modulatory Therapeutic Combinations in Hematologic Malignancies
Principal Investigators: Brian J. Druker, M.D., Adam Margolin, Ph.D., and Jeffrey Tyner, Ph.D.
Chemotherapies and targeted therapies are a major focus of drug development for acute myeloid leukemia (AML) and chronic lymphocytic leukemia (CLL), but the majority of patients eventually develop resistance to these treatments. Hence, there is a need to better understand the pathways underlying drug resistance and identify novel drugs or combinations of drugs that can effectively inhibit these pathways. The CTD2 Center at Oregon Health and Science University (OHSU-1) aims to take advantage of computational, experimental, and clinical expertise to infer mechanisms underlying drug resistance in AML and CLL and understand the pathways to predict effective drug combinations.
Through Beat AML and other programs, OHSU-1 has amassed a large cohort of patient samples with genomic, functional, clinical, and immune annotation. The Center plans to use this unique and large dataset to elucidate the molecular processes underlying drug sensitivity and resistance in AML and CLL patients. To achieve this goal, OHSU’s CTD2 Center proposes to i. develop an integrated computational framework, Predictors of Cellular Phenotypes to guide Therapeutic Strategies (PRECEPTS), ii. create a discovery resource of 400 leukemia patient samples with genomic and immune profiling and ex vivo genome-wide functional CRISPR/Cas screens, and iii. validate the predicted targets/biomarkers and test combinations of therapies to minimize or eliminate drug resistance by ex vivo testing. The proposed studies will have direct translational relevance in selecting novel treatment strategies for clinical trials and will be of benefit to other CTD2 Centers as well as the scientific community at large.
Integrative Bioinformatics and Functional Characterization of Oncogenic Driver Aberrations in Cancer
Principal Investigator: Gordon Mills, M.D., Ph.D.
Large-scale national and international cancer genomic studies are generating a compendium of tumor associated genomic alterations. Prioritizing these alterations as the most promising therapeutic targets for drug development is a major challenge. Although much is known about the function and clinical impact of recurrent “hotspot” aberrations in well-known cancer genes, less is known about which and how the more abundant, low-frequency mutations contribute to tumor progression. Evaluation of low-frequency alterations is difficult as they may indirectly influence tumor progression by modifying activities of concurrent driver aberrations. Differentiating between driver vs passenger gene alterations is also tricky as the driver activity is determined by the context of a given cancer. Hence, translation of tumor genomic datasets into effective cancer therapeutics will require new experimental systems to inform the functional activity of aberrations in the relevant biological context encompassing inter- and intra-tumoral heterogeneity. The CTD2 Center based at the Oregon Health and Science University (OHSU-2) will use state-of-the-art, high-throughput informatic and experimental approaches to functionalize oncogenic driver genes and determine the effects of single and combination drug therapies on tumor heterogeneity, as well as elucidate underlying mechanisms and therapeutic liabilities generated by driver aberrations.
In the preceding phase, the Center developed high-throughput gene cloning platforms and engineered numerous mutant and fusion genes for functional evaluation and delivery. In the current phase, OHSU-2 proposes to i. implement an algorithmic framework for prediction of oncogenic, gain-of-function driver aberrations of glioblastoma multiforme, pancreatic ductal adenocarcinoma, and epithelial ovarian cancer, ii. execute context-specific functional screens for the selection of single and combinatorial gene drivers of tumor progression and determine the effects of single and combination drug therapies on tumor heterogeneity and drug resistance, and iii. elucidate underlying mechanisms and therapeutic liabilities generated by driver aberrations. These studies will improve the understanding of how cancer gene aberrations affect downstream function within the protein and cellular pathways in cancer.
Organoid-based Discovery of Oncogenic Drivers and Associated Transformation Mechanisms
Principal Investigators: Calvin J. Kuo, M.D., Ph.D., Hanlee Ji, M.D., and Christina Curtis, Ph.D.
Cancer has extremely complex origins with a multitude of genomic and epigenomic alterations in interaction with cell-extrinsic stromal and environmental factors. Large-scale cancer genomic studies have generated a torrent of multi-scale omics data spanning mutations, gene expression, epigenetics, and proteomics. The data has revealed a daunting, highly plastic cancer landscape with tremendous interpatient variation requiring precision medicine approaches. In turn, this has generated an acute need for scalable cancer models to functionally characterize putative oncogenic driver events in a context dependent manner and elucidate the molecular drivers of treatment response. The CTD2 Center at Stanford University aims to apply state-of-the-art systems biology and bioinformatics to scan large-scale cancer genomic characterization studies. These approaches provide guidance in nominating candidate genes for functional validation in cancer.
In the preceding phase, Stanford developed in vitro 3D “organoid” culture methods (air-liquid interface) for cancer modeling. Organoids are three-dimensional miniature organs retaining tissue architecture and multilineage differentiation and represent a potentially transformative approach for functional interrogation of large-scale cancer genomic/epigenomic data sets. In the current phase, the CTD2 Center plans to i. prioritize amplified or deleted copy number alterations “outliers” from The Cancer Genome Atlas study for functional validation in lung, colon and stomach organoids, ii. functionally validate hypomethylated oncogene candidates and identify loci exhibiting non-linear relationships between hypomethylation and overexpression, and iii. study tumor genome/epigenome evolution and origin of cooperating oncogenic events. The Center’s focus on post-genomic systematic and functional interrogations translate the genomic findings into clinical application.
A Rational Systematic Approach to Find Combinations of Pharmacologic and Immune Therapies that Target Identifiable Oncogenic States
Principal Investigators: Pablo Tamayo, Ph.D., Ezra Cohen, M.D., Dan S. Kaufman, M.D., Ph.D., and Jill P. Mesirov, Ph.D.
Tumors have biological and clinical heterogeneity, even when they share the same mutations. Standard cancer classification is based on the anatomic site of tumor origin or the mutation status of known drivers; whereas, oncogenic state is defined as a functional status that is activated by specific signaling pathways and provides a more rational basis to identify therapeutic interventions. In addition, the wide variability of clinical responses to immunotherapy, and the onset of immune escape, are becoming a formidable obstacle to fully realize the potential of many new and effective immunotherapies. The CTD2 Center at the University of California San Diego (UCSD) aims to identify oncogenic cell states and characterize the most salient genomic and immune hallmarks to infer optimal combinations of pharmacologic and immunological perturbagens.
The Center’s preliminary data suggested that in each identifiable oncogenic state there is a close interplay between activation of oncogenic elements, cellular pathways, and the immune microenvironment. To validate this data, the Center proposes to i. select 5-10 oncogenic states and identify state-specific immunological targets, including immune checkpoints, neoantigens/anti-tumor epitopes, antibodies, and chimeric antigen receptors for cellular immunotherapy, ii. develop a multifactorial predictive model for each oncogenic state to identify the most effective combinations of pharmacological and immunological perturbagens, and iii. validate these perturbagens in isogenic cell systems, cancer cell lines, genetically engineered mouse models, and patient-derived xenografts. These studies will lead to the development of novel treatment strategies and provide the foundation for a new generation of more comprehensive, functional-based, precision oncology approaches.
The Cancer Target Discovery and Development Network at UCSF
Principal Investigators: Michael McManus, Ph.D., Jonathan Weissman, Ph.D., Trevor Bivona, M.D., Ph.D., and Sourav Bandyopadhay, Ph.D.
The major challenges of developing effective cancer treatments are drug resistance, tumor heterogeneity, and tumor evolution. Molecular characterization studies have identified a large number of genetic alterations in many cancer types. However, how these alterations contribute to cancer is not completely understood; hence, systematic exploration of the functional roles of these alterations is critical for the development of effective cancer therapeutics. The CTD2 Center at the University of California San Francisco (UCSF-1) aims to combine in-depth mining of large-scale genomic data and systems biology analyses to characterize functional roles of genetic lesions, both alone and in combination, in driving tumor formation and growth.
In the preceding phase, UCSF-1 developed CRISPR screening and ultra-high-throughput single cell protocols to perform comprehensive quantitative screens and identify genes essential for cancer initiation, maintenance, and possibly metastasis. In the current phase, the Center proposes to i. utilize the novel single-cell CRISPR platform to functionalize the cancer genomic data and associate genes with novel drug resistance mechanisms, ii. organize recurrently altered cancer genes from genome characterization initiatives into pathways associated with clinical resistance to understand functional impacts of inter- and intra-tumoral heterogeneity, iii. develop and apply methodologies to annotate pathways critical for tumor microenvironment interactions, and iv. construct genetic epistasis maps to identify and distinguish cancer drivers vs. passengers to uncover the optimal combinations of perturbagens with the potential to eliminate all cancer cells, despite their clonal heterogeneity and environmental context.
Integrating Targeted and Immunotherapy to Treat Genetically Heterogeneous Cancers
Principal Investigators: William Weiss, M.D., Ph.D., Allan Balmain, Ph.D., and Matthew Krummel Ph.D.
Cancer immunotherapy has attracted enormous attention by the recent success of immune checkpoint blockade inhibitors, such as anti-cytotoxic T lymphocyte antigen-4 and the anti-programmed death-1 antibodies. However, only a fraction of patients responds to checkpoint inhibitors. A major challenge is to improve these response rates by understanding the variables that influence clinical outcomes. Combining immune modulatory drugs, or adding these to targeted agents or chemotherapies, may improve clinical responses. The goal of the CTD2 Center at the University of California San Francisco (UCSF-2) is to identify, validate, and integrate targeted therapy with immunotherapy that most efficiently attack both tumor cells and the immune components in advanced cancers.
In the preceding phase, the Center developed immunocompetent mouse models that represent different ends of the spectrum of tumor types. Carcinogen-induced squamous carcinomas with high mutation load, but variable responses to immunotherapy, are “hot” tumors that present more or stronger antigens, or that encourage infiltration by immune effector cells. On the other hand, genetically engineered neuroblastoma with very low mutation burden are immunologically “cold” tumors that do not engage the immune system. In the current phase, UCSF-2 aims to address mechanisms of immune escape by exploiting these unique mouse models that mirror major genetic categories of human cancer – high vs low mutation load and strong vs weak immune infiltrate. Ongoing experiments aim to make immunologically “cold” tumors into “hot” tumors. To achieve this goal, the Center proposes to i. perform CRISPR/Cas9 screens in immune cells (monocytes and T cells) of the tumor microenvironment to identify genes associated with immune cell entry and effector functions, ii. use genetic or pharmacological perturbation of existing and newly identified candidate genes to determine pathways that improve/drive antigen presentation and abundance, and iii. implement computational analysis to exploit gene expression networks of existing databases, to prioritize potential targets that enhance immune responses. These studies will improve immunotherapies and generate lead targets, reagents, and diagnostics for cancer treatment.
Preceding Phase
Identifying and Targeting Cancer Dependencies With Small Molecules
Principal Investigator: Stuart L. Schreiber, Ph.D.
The Broad CTD2 Center aims to accelerate the discovery of patient-matched therapeutics. Cancers, as a result of their specific genetic or cellular features, acquire dependencies required for survival. Some drugs have been developed to target these dependencies and can yield high clinical response rates. However, drugs in this category benefit only a small fraction of cancer patients and, due to drug resistance, the beneficial responses are not always durable. The Broad CTD2 Center is discovering cancer dependencies that are targetable with small molecules and identifying combinations of drugs that can avoid or overcome resistance.
As part of the CTD2 Network, the Broad Center generated an 'Informer Set' of small-molecule probes, drugs, and select combinations. These compounds selectively target distinct nodes in cell circuitry that collectively modulate a broad array of cell processes. The Center quantitatively measured the sensitivity of deeply characterized cancer-cell lines to the Informer Set compounds, and undertook analyses to connect the sensitivities to cancer-specific alterations, including mutations, gene expression, copy-number variation, and cellular features such as lineage. These analyses and links to the underlying data (e.g., raw or processed data, or fitted curves) are provided openly on the CTD2 Data Portal and on an interactive public resource called the Cancer Therapeutics Response Portal (CTRP).
CTRP is a living resource for the biomedical research community, meaning the Center will continue to add data and analyses to updated versions of this interactive resource. The expanded dataset underlying the latest versio of CTRP includes an Informer Set of 545 compounds and select combinations tested for sensitivity in 907 cancer cell lines (specific analyses may be available only for relevant subsets of these totals). It can be mined to develop insights into small-molecule mechanisms of action and novel therapeutic hypotheses, and to support future discovery of drugs matched to patients based on predictive biomarkers. Future versions of the CTRP will include data generated under the following aims:
Aim 1. Integrate small-molecule, RNAi, and overexpression data from genomic cancer cell line profiling within the interactive resource (CTRP) to enable hypothesis generation
Aim 2. Identify and test hypotheses suggested by the interactive resource about cancer genetic dependencies targeted by small molecules
Aim 3. Use the interactive resource to identify combinations of small-molecule agents targeting cancer genetic dependencies that avoid or overcome drug resistance seen with single agents
Computational and Functional Approaches to Validating Cancer Genome Targets
Principal Investigator: Scott Powers, Ph.D.
The Cold Spring Harbor Laboratory (CSHL) CTD2 Center examines the vast repertoire of alterations in cancer genomes to discover and functionally validate therapeutic targets. Using sophisticated bioinformatics and high-throughput biological tools, the Center predicts gene sets and networks that are significantly altered in tumors and then uses human and mouse models to annotate drivers, dependencies, and interactions that may be vulnerable to combinatorial targeting. Some tools and approaches used by the Center for identification and in-depth functional validation of targets include:
- Computational technology that uses drug response data to infer network models that predict cellular responses to perturbations
- High throughput cDNA screening to identify oncogenic drivers
- High throughput shRNA screening to discover tumor cell dependencies and combinatorial targets
- Three dimensional “organoid” cell culture system that more accurately recapitulates in vivo tissue composition and architecture
- Flexible and rapid “speedy mouse” technology to evaluate the function of single or multiple genes in parallel
To date, the Center has identified and validated over fifty driver genes and/or dependencies; several are compelling therapeutic targets, and two are being tested in the clinic. Fibroblast growth factor 19/fibroblast growth factor receptor 4 (FGF19/FGFR4) is being targeted in a clinical trial for liver cancer and bromodomain containing 4 (BRD4) is in a trial as a target specific to Acute Myeloid Leukemia. The experimental strategies of the CSHL Center provide a blue print that can be adapted by the research community to identify targets across different tumor types.
Systems Biology of Tumor Progression and Drug Resistance
Principal Investigator: Andrea Califano, Ph.D.
At Columbia University, CTD2 funding supports efforts to study the systems biology of tumor progression and drug resistance. Researchers led by principal investigator Andrea Califano have developed a pipeline called Cancer Target High-Throughput Optimized Discovery and Evaluation (caTHODE). This pipeline uses both computational and experimental methods to efficiently discover and validate master regulators within the genomic networks that give rise to specific cancer subtypes. Master regulators are key nodes within networks of interacting genes and proteins that act as bottlenecks through which many different cellular signals must pass to initiate downstream activity. For this reason, researchers believe that master regulators may constitute points of vulnerability within a tumor. By computationally predicting and then experimentally validating the roles of master regulators in tumor progression and resistance to chemotherapy, this work is helping to generate a genome-wide list of prioritized targets for further investigation.
The caTHODE pipeline, which is intended to be scalable and effective for any tumor phenotype, utilizes a combination of methods developed at Columbia University. These include:
- Computational algorithms developed in the Califano laboratory that dissect and interrogate networks of transcriptional, post-transcriptional, and post-translational regulatory interactions.
- Genome-wide RNAi screens developed in the laboratory of José Silva (Icahn School of Medicine at Mount Sinai) to validate these master regulators.
- High-throughput chemical screening assays developed in the lab of Brent Stockwell to identify and validate small-molecule inhibitors of targets associated with phenotypes for tumor progression and drug resistance.
To date, the Columbia Center’s contributions to CTD2 include the discovery and validation of therapeutic targets, chemical modulators, and biomarkers in three distinct tumor subtypes: glioblastoma multiforme; glucocorticoid resistant T cell acute lymphoblastic leukemia; and an aggressive subtype of diffuse large B cell lymphoma that originates from the progression of follicular lymphoma. In addition, Columbia researchers developed collaborations with other CTD2 Centers focusing on additional cancer subtypes. These studies enabled the following findings:
Glioblastoma multiforme: In the mesenchymal phenotype of glioblastoma, the Columbia Center identified four modulators that harbor mutations. Mesenchymal glioblastoma is associated with the worst clinical outcomes for patients with brain cancers. In collaboration with Stuart Schreiber (Broad Institute), they also identified several novel inhibitors of signal transducer and activator of transcription 3 (STAT3), a key transcription factor within the regulatory network that promotes the mesenchymal phenotype.
T cell acute lymphoblastic leukemia: Researchers at Columbia University identified several candidate master regulators of glucocorticoid (GC) resistance and validated three genes. When these genes were silenced, GC-induced apoptosis increased and GC transcriptional activity was activated. Biochemical and functional assays revealed a mechanism of glucocorticoid resistance, and high-throughput screening uncovered an experimental compound that restores GC sensitivity. In a follow-up study, they inferred and validated additional transcription factors that are master regulators of GC resistance.
Diffuse large B cell lymphoma (DLBCL): NF-κB pathway activation is a hallmark of the most aggressive form of DLBCL, the activated B cell DLBCL (ABC-DLBCL) subtype. The Columbia team identified transcription factors and signaling molecules that are critical to ABC-DLBCL and identified master regulators that contribute to follicular lymphoma transformation.
Ovarian serous cystadenocarcinoma: To identify molecular mechanisms of ovarian cancer pathogenesis, Columbia Center researchers reconstructed the transcriptional, post-transcriptional, and post-translational networks of ovarian serous cystadenocarcinoma. They identified master regulators that indicate poor prognosis, drive tumorigenesis, and promote resistance to cisplatin chemotherapy. Factors within regulatory networks that modulate the activity of the master regulators were also revealed.
Non-small cell lung cancer: To dissect the genome-wide signal transduction network that is regulated by tyrosine kinases, the Califano Lab applied a modified version of the Algorithm for the Reconstruction of Accurate Cellular Networks (ARACNe) algorithm (pARACNe) to a published dataset of phospho-proteomic profiles of non-small cell lung tumors. They also used a modified version of the Master Regulator Inference Algorithm (MARINa) algorithm to compare gene expression patterns and identify master regulators in 50 cell lines.
Functional Annotation of Cancer Genomes
Principal Investigator: William C. Hahn, M.D., Ph.D.
The comprehensive characterization of cancer genomes has and will continue to provide an increasingly complete catalog of genetic alterations in specific cancers. However, most epithelial cancers harbor hundreds of genetic alterations as a consequence of genomic instability. Therefore, the functional consequences of the majority of mutations remain unclear.
The Dana-Farber Cancer Institute (DFCI) CTD2 Center has developed tools to perform somatic cell genetics in mammalian cells. Tumor cell populations are systematically analyzed to identify the function of somatically altered genes identified by genome characterization studies. These approaches also reveal co-dependencies or synthetic lethal partners in tumors.
The Center is using high-throughput genetic perturbation approaches to create genome-wide datasets and applying bioinformatics to identify and credential targets. As part of this effort, the Center is cataloging all essential genes that contribute to proliferation or survival in a large number of ovarian and colon cancer cell lines. Novel bioinformatics approaches are being used to interrogate this functional data and integrate it with other genomic datasets. These approaches in vitro and in silico will identify targets (e.g., candidate oncogenes that are mutated or amplified and essential) in specific genetically defined tumor subtypes. Complementary to this approach, the Center is using multiplexed tumor formation assays and context-specific transformation assays to identify novel oncogenes. They have also developed methods to identify and validate targets in vivo in patient-derived experimental models. Beyond discovery, DFCI CTD2 investigators will apply high-throughput gene expression profiling and targeted proteomics to interrogate the signaling networks perturbed by these oncogenes.
All of the outputs of these investigations (data, methodologies and bioinformatics tools) are made readily available through the CTD2 Data Portal. Access to this information will enable the cancer research community to identify targets based on both genomic and functional biological evidence. Ultimately, these data will inform the most appropriate genetic context for downstream mechanistic and validation studies and prioritize targets for translation into therapeutics.
High Throughput Protein-Protein Interaction Interrogation in Cancer
Principal Investigator: Haian Fu, Ph.D.
Genomic alterations in various tumor types, as revealed by cancer genomics initiatives such as The Cancer Genome Atlas (TCGA), often lead to re-wired protein-protein interaction networks, which in turn drive tumor initiation and progression. Thus, identifying prominent PPI nodes and networks among oncoproteins and tumor suppressors as enriched by various genomics datasets and the validation of their critical roles in tumorigenesis and progression are expected to reveal an entirely new class of PPI-based cancer targets for therapeutic development.
The Emory Molecular Interaction Center for Functional Genomics (MicFG), with its expertise in high throughput technologies for the study of protein-protein interactions (PPI), productive track record of innovative HTS assay development for chemical lead discovery, and proven cancer genomics mining, database, and data integration capabilities, proposes to utilize high throughput PPI technologies to interrogate cancer genomes through a team science approach (i) to rapidly establish oncogenic PPI networks of selected cancer types based on TCGA and other genomic datasets, (ii) to validate functional roles of key PPI nodes or hubs in tumorigenesis and progression, (iii) to develop HTS assays for critical tumor-associated PPIs to enrich the therapeutic target pipeline of the NCI and the drug discovery field, and (iv) to leverage our informatics capability for genomics data mining for prioritized PPI mapping, functional PPI validation, and trans-network data sharing and collaboration.
We aim to bridge the gap between vast genomic datasets and therapeutic discovery by establishing and interrogating the cancer PPI target space. With our PPI expertise and state-of-the-art high throughput technologies, which are highly adaptable to a variety of outputs, we intend to function as an active, synergistic member in collaborative trans- Network projects.
An Integrated Functional and Computational Discovery Engine for Preclinically-Validated Cancer Drug Targets
Principal Investigator: Christopher Kemp, Ph.D.
Large-scale molecular analyses have provided an unprecedented global view of the molecular defects in cancers and promise to revolutionize precision cancer medicine by guiding the development of therapies that are matched to genomic alterations in tumors. However, developing and implementing successful targeted therapies remains a daunting challenge. Cancer genomes are complex, so determining which genes to target from the hundreds of possibilities is challenging. Many of the best known oncogenes and tumor suppressor genes have not been successfully targeted directly. In cases where targeted therapies are used clinically, successes have been limited because patients frequently develop drug resistance, underscoring the need for combination therapies. The Fred Hutchison Cancer Research Center (FHCRC) CTD2 Center and collaborators at Oregon Health and Science University have developed technology and methods to help address these challenges.
The FHCRC CTD2 Center performs genome-scale functional testing of potentially druggable genes, including candidates derived from TCGA and other large genomic datasets, using isogenic cell lines and genomically characterized patient-derived tumor cell cultures. They use cell viability and other assays as readouts for high-throughput well-based siRNA and therapeutic compound screens. Integrating the results of these functional genomic approaches with genotype-specific vulnerabilities, which are inferred through computational analyses of large-scale public datasets, enables the identification of genotype- and patient- specific therapeutic targets that are selectively lethal to human cancer cells carrying defined mutations.
Currently, the Center uses these strategies to identify and credential novel drug targets for cancers most in need of better therapies, including aggressive subtypes of head and neck squamous cell carcinoma, pancreatic ductal adenocarcinoma, and triple negative breast cancer. Gene targets that are identified and have known pharmacologic inhibitors are tested alone and in combination with existing standard of care drugs to nominate candidate targets for preclinical validation or clinical trials.
Identifying and Validating Tumor-Selective Targets for use in Immunotherapy
Principal Investigators: Martin McIntosh, Ph.D. and Edus H. Warren, M.D.
The CTD2 Center at Fred Hutchison Cancer Research Center (FHCRC) aims to identify novel cancer-specific antigenic targets for immunotherapeutic approaches that are specifically toxic to patients’ malignant cells. The Center is identifying numerous promising cancer-specific targets in lung adenocarcinoma and serous ovarian carcinoma. Development of treatments to selectively attack cancer cells requires identification of cell surface proteins or protein complexes that have three essential elements: abundant on malignant cells, absent or rare in somatic tissues, and recognized specifically by a T cell or B cell receptor. The FHCRC Center has developed a high-throughput pipeline to efficiently identify cancer-specific targets that satisfy all three elements and are naturally suited for immunotherapeutic approaches because of their potential to be targeted without adversely affecting normal tissues.
FHCRC identifies potentially immunogenic peptides that arise from cancer-specific changes in a protein’s amino acid composition and are recognized by human T cell or B cell receptors. This recognition activates the body’s immune system to designate the tumor as foreign material and initiate a response to destroy it. Targets for T cell based immunotherapy must be on the cell surface, either as a peptide that is presented by major histocompatibility complex (MHC) class I protein complexes or as a cell surface protein variant. To discover potential candidate targets, the Center applies computational methods to RNA-seq data from The Cancer Genome Atlas and the Genotype Tissue Expression project. These data are combined with FHCRC/University of Washington Comprehensive Cancer Center data that profiles translating ribosomes. Variant coding transcripts that undergo a cancer-specific splicing event and are common in tumors, but are rare or not found in normal tissues, are designated cancer-specific transcripts (CSTs). CSTs that are bound to translating ribosomes or associated with the endoplasmic reticulum (found by another type of RNA-seq assay) are then used to predict the cancer-specific polypeptides (CSPs). These CSPs are experimentally tested to determine if they harbor peptides presented by the MHC complex or if their variant is presented on the cell surface. All discovery and verification efforts use materials from patients who participate in a FHCRC/University of Washington Comprehensive Cancer Center study.
The CSPs are developed for use in adoptive T cell therapies. To start therapy development, the team identifies from patients or unaffected individuals a human T cell that recognizes the MHC-bound antigen, or a B cell that recognizes the cell surface proteins The T cell or B cell receptor sequence (TCR and BCR, respectively) is then obtained and used to engineer the therapeutic T cell. If necessary, the TCR may be modified to have higher affinity to the antigen. BCR sequences are used to engineer a T cell that adopts the recognition capability of an antibody. This is commonly called Chimeric Antigen Receptor T cell (or CART) therapy. Before they are tested as therapeutics in clinical trials, the active T cells are tested in vitro to ensure the modified receptors recognize the cancer-specific target and in vivo to determine if they elicit the desired immune response. When these methods are used in a patient, his or her T cells are harvested, engineered to express the specific TCR or BCR, and possibly other receptors, and then re-infused into the patient.
The FHCRC Center has active collaborations within and outside the CTD2 Network. Other CTD2 investigators exploit cancer-specific cell surface proteins identified by FHCRC to deliver toxic materials specifically to malignant cells. To validate targets in biomaterials collected from patients, the Center collaborates with leading clinical researchers who focus on moving therapies from mouse models into human trials.
Functional Analysis of Oncogenic Networks in Primary Organoids
Principal Investigators: Calvin J. Kuo, M.D., Ph.D. and Hanlee Ji, M.D.
Cancer arises from the acquisition and concerted action of multiple mutations and genomic aberrations in discrete combinations of tumor suppressors and oncogenes, known as "drivers". Large cancer genome-scale sequencing studies such as TCGA, are now operative and the Cancer Target Discovery and Development (CTD2) Network seeks "to bridge the gap between the enormous volumes of data generated... and the ability to use these data for the development of human cancer therapeutics". A secondary goal for the CTD2 Initiative is that in five years, "the entire CTD2 Network is expected to identify and characterize targets for approximately 25 or more (if possible) cancer types," and for applicants "to have or build the capacity for in depth analyses and experimental approaches utilizing datasets for many cancer types." A broad "coverage" is the paradigm for this initiative.
The wealth of TCGA data will be directly coupled to robust in vitro functional validation of candidate cancer driver modules using primary mouse 3D organoid cultures of diverse tissues arrayed in high-throughput format. In Aim 1, the Hanlee Ji and Sylvia Plevritis groups will identify co-segregating mutational modules from TCGA datasets from multiple solid tumor types, using complementary methods of supervised Bayesian analysis and Unsupervised Module Network Analysis for Master Regulators. In Aim 2, these prioritized mutational modules, stratified for clinical significance, will undergo direct functional validation in a broadly applicable, multiplexable, in vitro 3D primary organoid system developed by the Calvin Kuo group, which is amenable to combinatorial gene engineering. In collaboration with Bill Hahn, this will utilize high throughput lentiviral introduction of cDNA or shRNA to systematically interrogate the genes within amplicons and deletions, contextually modeled in the TCGA mutational background in which these copy number variations occur. Additionally, co-segregating mutational modules from diverse tissues will undergo systematic deletion in organoid cultures to define minimal module composition, and we will pursue process development to extend the range of tissues from which organoids can be modeled.
Overall, these studies describe bioinformatic and in vitro modeling approaches that are robustly portable across a variety of organ systems for functional interrogation of diverse TCGA datasets and with attendant implications for cancer biology, diagnosis and therapy.
Systematic Development of Novel Druggable Targets in Glioblastoma
Principal Investigator: Michael E. Berens, Ph.D.
Gene expression patterns across glioblastoma (GBM) cases in The Cancer Genome Atlas (TCGA) were previously queried by a novel computational analysis tool for mining molecular “contexts”. The analysis uncovered 12 distinct gene-based molecular “contexts of GBM”. Based on molecular profiles from previously established and clinically relevant preclinical models, the Translational Genomics Research Institute (TGen) CTD2 Center assembled a collection of 54 patient-derived xenograft GBM models that reside within the same genomic-based contexts. This collection represents diverse GBM tumor phenotypes. Collaborators at TGen, Sanford Burnham, and Thomson-Reuters use these PDX models, in combination with bioinformatic and empiric approaches, to achieve the following goals:
- Identify novel targets associated with subclasses of GBM and discrete features of the “hallmarks of cancer”
- Functionally validate prioritized targets in vitro using high throughput RNAi and chemical based assays
- Validate prioritized targets in vivo using orthotopic human GBM tumor grafts and inducible shRNA and/or hits from chemical based screening assays
Within each distinct context, bioinformatic algorithms (MetaBaseTM) query the patterns of expressed genes and the pathways in which they function. The known networks in which these genes reside are used to discern candidate therapeutic targets specific for certain GBM contexts. Chemical biology libraries and RNAi panels (well characterized for mechanism of action) uncover molecules that specifically and uniquely influence GBM cell behaviors within certain contexts. Compounds identified in empiric studies are also used in high throughput screening against short-term cultures from the GBM PDX models. Each of these strategies can be adapted to assess various hallmarks of cancer, including proliferation, survival, self-renewal, migration, invasion, and apoptosis-resistance.
Integrating in silico and laboratory technologies continuously improves computational approaches to help prioritize targets in the specific tumors. Furthermore, small-molecule screens against identified targets and pathways provide a potential fast-track to develop novel “perturbagens” against the validated targets. Sub-classifying GBM during these experimental processes informs fundamental insight into disease biology and lays the foundation for precision-guided therapy for molecularly-subgrouped human cancers.
By demonstrating an efficient approach for tractable target identification and validation, this project will impact the fields of informatics, cancer biology and drug discovery and accelerate, the translation of genomic discoveries into new treatments.
Transformative Strategies for Dissecting Cancer Pathways
Principal Investigators: Frank P. McCormick, Ph.D., Michael T. McManus, Ph.D. and Jonathan S. Weissman, Ph.D.
The ability to effectively and efficiently perturb endogenous gene expression is essential to uncover tumor-specific vulnerabilities that may be targeted with therapeutics. However, targeting single vulnerabilities in patients often leads to drug resistance and treatment failure. Therefore, the signaling pathways that act synergistically to promote tumor growth must be identified to design effective combination cancer therapies (polytherapies) that target key cancer "driver" pathways. The lack of a systematic method by which to identify tumor-specific vulnerabilities in pathways that functionally cooperate to drive tumor growth creates a major challenge in developing such therapies. Therefore, the search for effective cancer polytherapies has been done largely in an ad hoc manner by exploring limited numbers of potential combinations. The CTD2 Center at the University of California San Francisco (UCSF-1) has developed an experimental pipeline with high-throughput technologies to systematically identify pathways that when targeted, lead to specific and synergistic destruction of cancer cells.
The UCSF-1 Center uses the “EXPAND RNAi” high-throughput screening approach based on complex RNAi libraries to identify candidate driver genes. To evaluate the relevance of the potential drivers, the Center quantifies responses to targeted inhibitors using screens in engineered primary cell lines (isogenic cell lines). The quantitative data are integrated with known information for each compound and cell line used in the screen to create chemical-genetic interaction maps. The maps provide critical insights into biological pathways and functional dependencies and can be used to identify and design potential polytherapies.
The UCSF-1 Center also employs a complementary approach to study gene function in cancer cells, the clustered regularly interspaced short palindromic repeats (CRISPR) technique. They modified the CRISPR system to dynamically repress gene transcripts (CRISPRi) as well as established a novel system that results in gene activation (CRISPRa). These methods exhibit low off-target effects and specifically target transcriptional start sites allowing the Center to apply the technology on a whole-genome scale.
Novel approaches developed by the UCSF-1 Center establish systematic methods to uncover gene interaction networks that drive tumor growth. The Center’s experimental pipelines will facilitate the development of cancer polytherapies and create new paradigms for discovering cancer therapeutics that fully capitalize on the genomic profiling of human tumors.
Genetic Network Analysis to Discover Cancer Targets
Principal Investigator: William A. Weiss, M.D., Ph.D.
Cancers acquire genetic changes, or drivers, that are required for tumor survival. Therapies developed to target these cancer drivers often lead to clinical responses. However, the promise of targeted therapies has been tempered in many cancer patients by eventual relapse due to therapy-driven drug resistance that results from rewiring of the tumor’s genetic architecture. Novel approaches are necessary to understand this rewiring, and identify upstream or downstream targets that may yield to small molecule modulation or other therapeutic strategies.
The CTD2 Center at University of California San Francisco (UCSF-2) has developed computational network tools to visualize the genetic architecture of tissue samples. These tools correlate gene expression patterns from heterogeneous samples of normal or tumor tissue. This approach identifies genetic networks that control tissue organization or cellular function in complex normal tissues, and details the rewiring of these networks during tumor evolution in vivo.
These methods were initially developed to analyze normal and tumor tissues from mouse skin. Transcriptional profiles of normal skin samples from wild type mice and mice lacking the tumor oncogene, Hras, were compared. This analysis provided insight on changes in genetic architecture that may lead to susceptibility to tumor development in tissues lacking Hras. Recently, these analyses have been extended to neural tissues from mouse models of neuroblastoma. If appropriate mouse models are available to generate expression profiling data on tissues of interest, the computational approach has the potential to identify the genetic architecture of tumor types being studied by other CTD2 Centers through trans-Network collaborations.
The UCSF-2 CTD2 Center is currently identifying candidate drug targets by mining genomic data from human neuroblastomas and carrying out functional shRNA screens in human models. Results from human data mining are compared with architecture analyses results from the neuroblastoma mouse model to prioritize targets for further investigation. Prioritized targets are analyzed computationally using gene expression and copy number data from multiple developmental stages of carcinoma including benign, malignant, and metastatic samples. This analysis will investigate stage-specific activation/inhibition of candidates and identify changes in their expression architecture during tumor progression. Targets will then be validated using gain- or loss- of function approaches in human derived cell lines and patient derived xenograft models from appropriate human cancers. Potential new targets will be tested for sensitivity to pharmaceuticals. These approaches will identify both new cancer targets and novel roles for existing therapeutics within cancer pathways.
Biological Annotation of Data from Large-Scale Cancer Genomic Initiatives
Principal Investigator: Gordon B. Mills, M.D., Ph.D.
Large-scale tumor profiling efforts by consortia such as The Cancer Genome Atlas (TCGA), Therapeutically Applicable Research to Generate Effective Treatments (TARGET) and the Cancer Genome Characterization Initiative (CGCI) are cataloging genomic aberrations across major cancer lineages. These studies have revealed an extraordinary level of genome complexity. The CTD2 Center based at the University of Texas MD Anderson Cancer Center (MDACC) is working to distinguish key “driver” events critical to pathogenesis from the numerous biologically-neutral “passenger” aberrations that accompany unstable tumor genomes. The Center’s goal is to find ways to validate functional driver aberrations, since targeting such events or their activated pathways may improve patient outcomes.
Currently, the CTD2 Center at MDACC examines the functionality of thousands of potential driver genes found within breast, pancreas, melanoma and endometrial tumors. MDACC, in collaboration with Baylor College of Medicine, has developed pipelines for the rapid construction of barcoded cDNAs with mutations identified through tumor profiling by the large-scale genomics consortia mentioned above. The Center is actively constructing every somatic event within candidate cancer genes because each change within a given gene may result in a different functional impact or therapeutic response. The cDNAs are suitable for expression in cell and animal cancer models.
To maximize discovery potential, cell lines expressing wild type or mutant cDNA can be used in multiple high-throughput screens in parallel. For example, generalizable sensor cell assays quantify the ability of the mutant genes to induce cell survival and proliferation. Through these assays, driver genes are identified. Cells expressing confirmed driver genes are subjected to compound screening platforms designed to identify drugs that inhibit driver activity. Together, these in vitro approaches permit broad evaluation of driver candidates across multiple cancer lineages. Importantly, these approaches complement RNAi-based gene depletion screening strategies for driver discovery used by other CTD2 Centers.
While cell-based screening systems are amenable to high-throughput analyses, in vitro models do not fully recapitulate all hallmarks of tumorigenesis and metastasis. To address this challenge, the Center has developed a Context-Specific Screen (CSS) platform to interrogate the tumorigenic potency of candidate driver genes under the appropriate in vivo genetic and microenvironment contexts. For in vivo screens, non-transformed human primary cells expressing pools of confirmed barcoded driver genes are implanted orthotopically at defined sites in the mouse to ensure the correct microenvironment context. Upon tumor formation, genes driving tumor progression are identified from tissues by barcode amplification and sequencing. Importantly, this approach permits high-throughput discovery of cooperating driver events that are co-selected in output tumors.
In summary, this work will provide the greater research community multi-level functional assessments of oncogenomics data collected by TCGA and other large scale genomic studies. This technology, which facilitates biological annotations of genomic changes will create unique opportunities for transformative cancer research by engaging the research community. Ultimately, these contributions will accelerate drug development and implementation.
A Concerted Attack on Patient-Specific Oncogenic Vulnerabilities in Lung Cancer
Principal Investigators: Michael Roth, Ph.D., Michael A. White, Ph.D., John B. MacMillan, Ph.D., and John D. Minna, M.D.
Lung cancer is a major cause of death in the United States and there is a clear need for additional and better therapeutic approaches. The multiple oncogenotypes that cause lung cancer are known to confer different responses to currently approved therapeutics. However, the molecular basis of these differences is not completely understood and a systematic approach to identify functional differences in cells derived from tumors with different oncogenotypes has not been carried out. The CTD2 Center at University of Texas Southwestern (UTSW) Medical Center uses two different high throughput approaches to address this need.
Using siRNA libraries, the Center studies how loss of function of each human gene affects survival of lung cancer cell lines that represent at least 6 different oncogenic subtypes. This high throughput technique measures cancer cell survival. Sensitivity to loss of a gene indicates that the gene (or another protein in its pathway) may be a potential therapeutic target. The targets identified are then tested for therapeutic potential in appropriate tumor models in animals.
The UTSW Center is also conducting high-throughput screens to identify compounds that may have therapeutic benefit in one or more subsets of genetically distinct lung cancers. In a pilot effort, the Center used a library of 200,000 synthetic drug-like compounds to screen for inhibition of cell survival in the same lung cancer cell lines used in the siRNA screens. The Center is now screening 40,000 natural products derived from marine bacteria. Compounds that inhibit cell survival in culture are tested for the ability to inhibit growth of human tumor explants. These experiments provide a potential fast-track for discovering novel therapies as well as a method for detecting tumor-specific vulnerabilities that are not detected by the RNAi strategy described above.
After identifying tumor specific vulnerabilities through the two independent screening methods, gene knock-down data and compound sensitivity data can be correlated to identify novel drug targets and compounds that inhibit them. A database of the combined functional genomic and compound screening results are available to the public through the CTD2 Data Portal and compound structures annotated with biological data are displayed on PubChem.