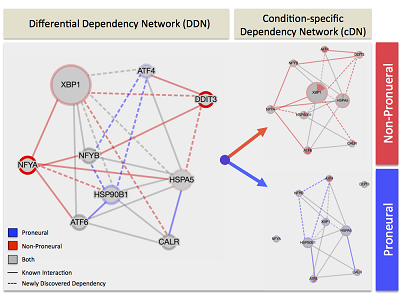
EDDY incorporates prior knowledge of gene interactions to generate interaction networks. Image courtesy of the author, Dr. Speyer.
We have previously developed a statistical method to identify gene sets enriched with condition-specific genetic dependencies. The method constructs gene dependency networks from bootstrapped samples in one condition and computes the divergence between distributions of network likelihood scores from different conditions. It was shown to be capable of sensitive and specific identification of pathways with phenotype-specific dysregulation, i.e., rewiring of dependencies between genes in different conditions. We now present an extension of the method by incorporating prior knowledge into the inference of networks. The degree of prior knowledge incorporation has substantial effect on the sensitivity of the method, as the data is the source of condition specificity while prior knowledge incorporation can provide additional support for dependencies that are only partially supported by the data. Use of prior knowledge also significantly improved the interpretability of the results. Further analysis of topological characteristics of gene differential dependency networks provides a new approach to identify genes that could play important roles in biological signaling in a specific condition, hence, promising targets customized to a specific condition. Through analysis of TCGA glioblastoma multiforme data, we demonstrate the method can identify not only potentially promising targets but also underlying biology for new targets. (Publication Abstract)
